How does the Common Mechanism flag sequences?
The Common Mechanism screens nucleotide sequences of 50 base pairs or more through the following screening steps:
- Biorisk search: Sensitively identify well-established sequences of concern, such as toxins and virulence factors. (This is done using a fast HMM-based search against sequence profiles curated from annotations of regulated pathogens and toxins.)
- Taxonomy Search: Query large protein and DNA databases to retrieve the organism the genome most closely matched to the query sequence, then cross-referencing that match with a variety of control lists, including those from India, China, and South Africa.
- Low-concern search: Clear earlier flags based on matches to common or conserved sequences. This database includes protein sequences found in thousands of bacterial species, RNA sequences that participate in processes essential for life, and sequences submitted to the iGEM parts registry with no associated safety flags.
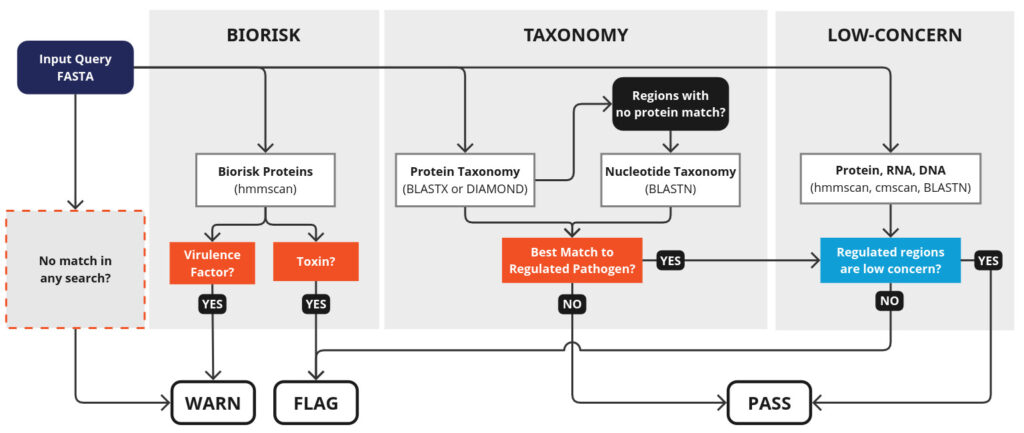
These steps, and how they lead to decisions to flag a sequence, are shown below: